
RAS Biochemistry and Biophysics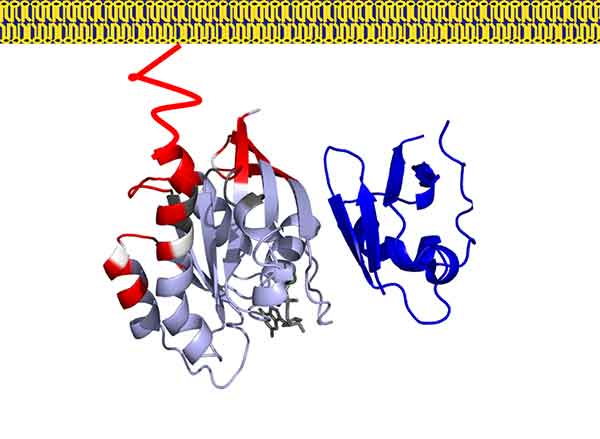
Model of RAS-RAF complex at the membrane
Credit: Que Van, NCI RAS Initiative
RAS proteins function through interactions with the inner plasma membrane, and with numerous interacting partner proteins. These interactions are not understood at atomic levels, and could hold the key to controlling RAS signaling in cancers. The goal of the Biochemistry and Biophysics group is to apply the most sensitive techniques to better understand how these interactions affect signaling by wild-type and mutant RAS. Our ProgressOur group has characterized the interactions of fully modified KRAS4b protein with nanodiscs of different lipid compositions. Nanodiscs are a surrogate for cell membranes where the lipid composition can be completely specified. In addition, protein-nanodisc complexes are amenable to biophysical assays. Only fully modified KRAS protein can bind to nanodiscs. Half a dozen different mutants of KRAS account for more than one million cancer deaths each year world-wide. Understanding the different mutant proteins in detail may help us understand why different KRAS mutants predominate in different cancers. Our group has measured the binding affinities and intrinsic hydrolysis rates of all the major oncogenic mutants of KRAS4b. We have utilized opportunities to collaborate on research, including the characterization of KRAS proteins by NMR, analytical ultracentrifugation, neutron scattering, and single molecule analysis in artificial lipid bilayers. View the PubMed abstract on the characterization of fully modified KRAS4b protein. Our Projects
Tools We Use
CollaborationsThe RAS Biochemistry and Biophysics Group has collaborated with: Paul Henderson Jay Groves Steve Sligar Marco Tonelli Rodolfo Ghirlando Frank Heinrich Carla Mattos Michael Gross Chris Stanley Contact
Dr. Andy Stephen, Biochemistry and Biophysics Group Lead For more information, contact the RAS Biochemistry and Biophysics Group team lead: Dr. Andy Stephen Updated: September 15, 2016 Syndicated Content Details:
Source URL: https://www.cancer.gov/publishedcontent/syndication/1061553.htm Source Agency: National Cancer Institute (NCI) Captured Date: 2016-09-13 19:10:23.0
|